These clients will allow you to discover and access data and analysis services provided through the BioMOBY framework. Some also allow the construction and execution of re-usable workflows and analytical pipelines.
| Client Title | Source | Platform & Language | Description |
| Gbrowse_moby | Mark Wilkinson | Web-based, cross-platform | The first Moby client ever written; very simplistic and limited in power. |
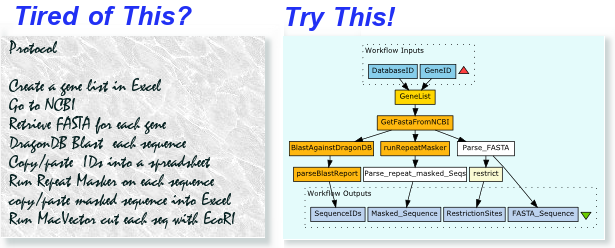
| Taverna | Tom Oinn, Martin Senger; myGrid | Java, cross-platform | Taverna allows linking of inputs and outputs from both Moby and non-Moby services into extensive and complex workflows. |
| MOWserv | Instituto Nacional de Bioinformatica | Web based, cross-platform | MOWserv allows construction and execution of workflows through a web interface |
| Remora | Genopole Toulouse | Web based, cross-platform | Remora allows design of Moby workflows through a web interface |
| Ahab | Benjamin Good, Clarence Kwan, Wilkinson Laboratory, UBC | Web based, cross-platform | Ahab allows parallel execution of multiple services simulteneously. |
| Seahawk | Paul Gordon, Sun Centre of Excellence, University of Calgary, Alberta, Canada | Java applet, cross-platform | Allows a user to load text or HTML data sources, then discover and execute MOBY Services through hyperlinks and text highlighting. |
Clients with embedded MOBY functionality
These are applications that provide a variety of bioinformatics functionality and take advantage of BioMOBY to extend that functionality.
| BioTrawler | Frank Gibbons, Harvard | Web based, cross-platform | Explore protein interaction networks by surfing MOBY! |
| BlueJay | Paul Gordon, University of Calgary (Genome Canada/Genome Alberta) | Web based, cross-platform | Explore genomes with MOBY out-linking functionality |
| BioFloWeb | Sophie Durand, INRA, France & PlaNet Consortium | Web based, cross-platform | Arabidopsis ‘gene cards’ via PlaNet MOBY services |
| AtiDB Client | Sean Walsh, John Innes Centre, UK | Web based, cross-platform | Arabidopsis Locus Report via PlaNet MOBY services |